library(FITSio)
library(SpectralUnmix)
X <- readFITS("cube.fits")
Mat <- cube_to_matrix(X)
Mat[!is.finite(Mat)] <- 0
Mat <- pmax(Mat, 0)IFU Workflow
FITS to matrix
Fit and inspect
fit <- spectral_unmix(Mat, k = 5, lr = 0.01, niter = 5000)
plot(fit, type = "spectra")
plot(fit, type = "maps", nx = dim(X$imDat)[1], ny = dim(X$imDat)[2])
plot_reconstruction(fit, Mat, n = 6)Example spatial maps from the bundled synthetic cube:
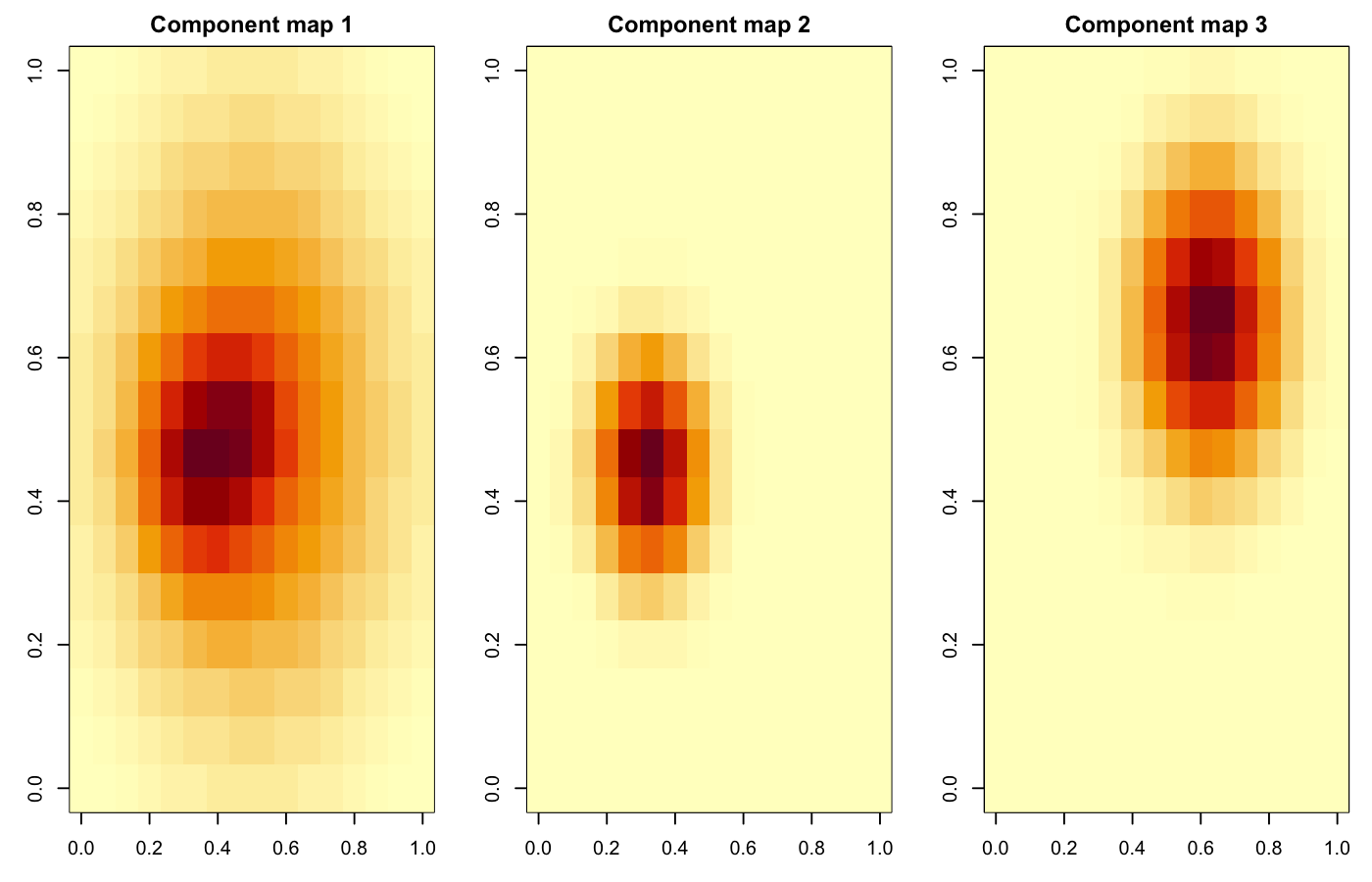
Reconstruction outputs
maps <- predict(fit, type = "spatial")
cube_hat <- predict(
fit,
type = "cube",
nx = dim(X$imDat)[1],
ny = dim(X$imDat)[2]
)
resid <- residuals(
fit,
x = Mat,
nx = dim(X$imDat)[1],
ny = dim(X$imDat)[2]
)Metadata and redshift
When the input to cube_to_matrix() is a FITS-like list object, non-image entries are carried forward as metadata. That metadata can be recovered from the fitted object or reconstructed cube with cube_metadata().
fit <- spectral_unmix(Mat, k = 5, lr = 0.01, niter = 5000)
cube_metadata(fit)
cube_metadata(cube_hat)If redshift information is stored in the FITS-side metadata, it will now be preserved as metadata and can be carried into fitted and reconstructed outputs.