Capivara 
Overview
Capivara provides spectral segmentation tools for Integral Field Unit (IFU) data cubes. Version 0.3.0 adds built-in missing-data support in the exact workflow, sparse-Ward segmentation for large cubes, Sagui-style white-light starlet masking, variance-aware spectral summaries, and SNR-guided component selection.
The core segmentation API is intentionally small:
segment()for the standard exact Ward workflow, including missing spectral channels.segment_large()for large cubes where exact pairwise distances are too expensive in RAM.
Current Capivara Mosaic
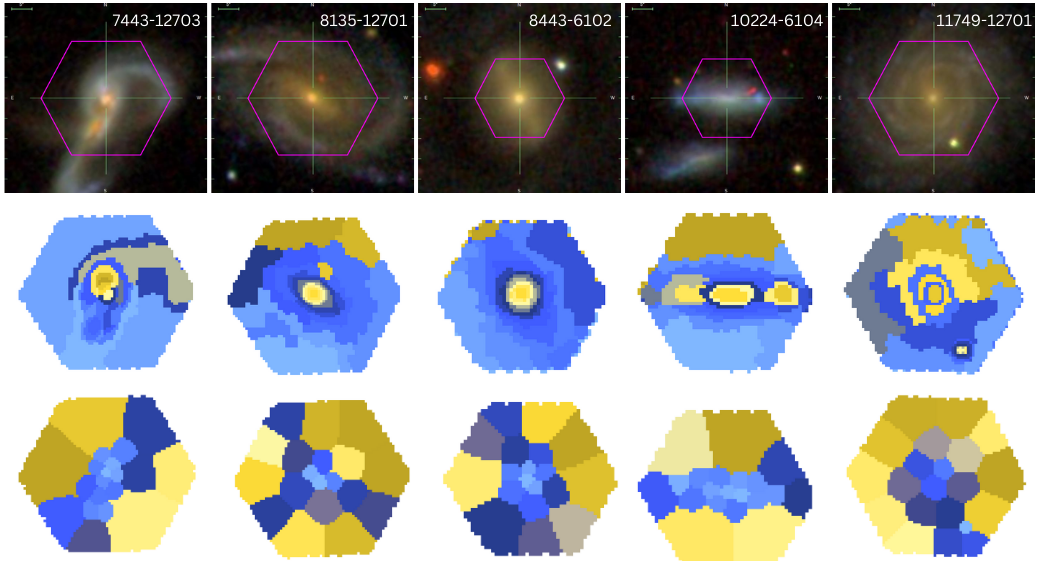
Sagui Comparison Mosaic
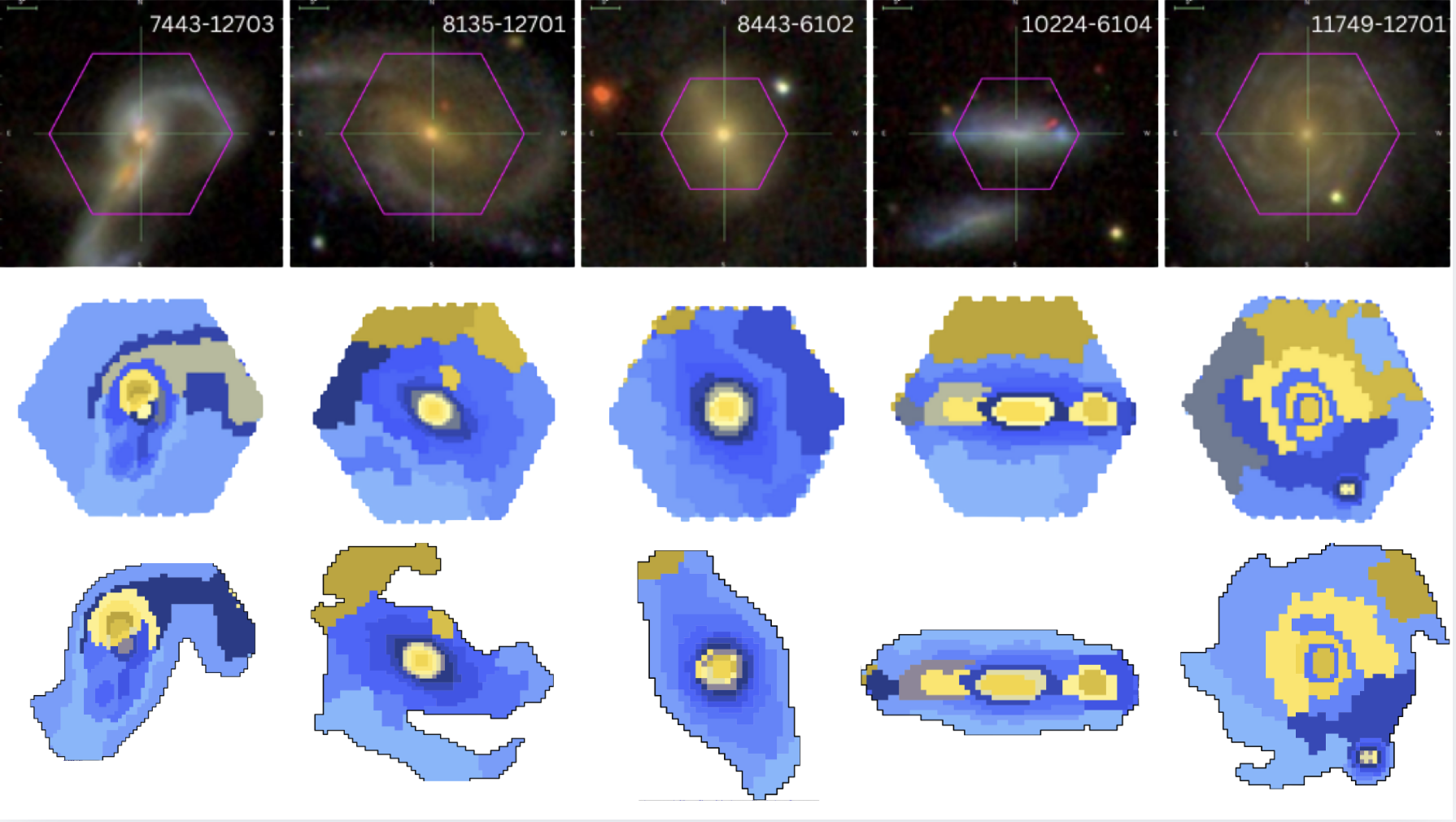
Both mosaics are displayed with the same fixed size to make the visual comparison easier.
What’s New In 0.3.0
segment()now handles missing spectral channels by default.segment_large()adds the calibrated kNN-restricted sparse-Ward backend for large cubes.estimate_segment_memory()reports the exact Ward distance-vector RAM lower bound before you allocate it.- both segmentation backends can optionally use a Sagui-style starlet mask from the white-light image.
summarize_cluster_spectra()supports median, summed, and weighted spectra.choose_ncomp_by_snr()selectsNcompfrom a variance-aware SNR cut.torchis now optional.
Installation
Optional GPU acceleration:
Usage
Basic Segmentation
Missing-data-safe Segmentation
segment() now handles missing spectral channels directly.
Large-cube Segmentation
Sagui-style Starlet Segmentation
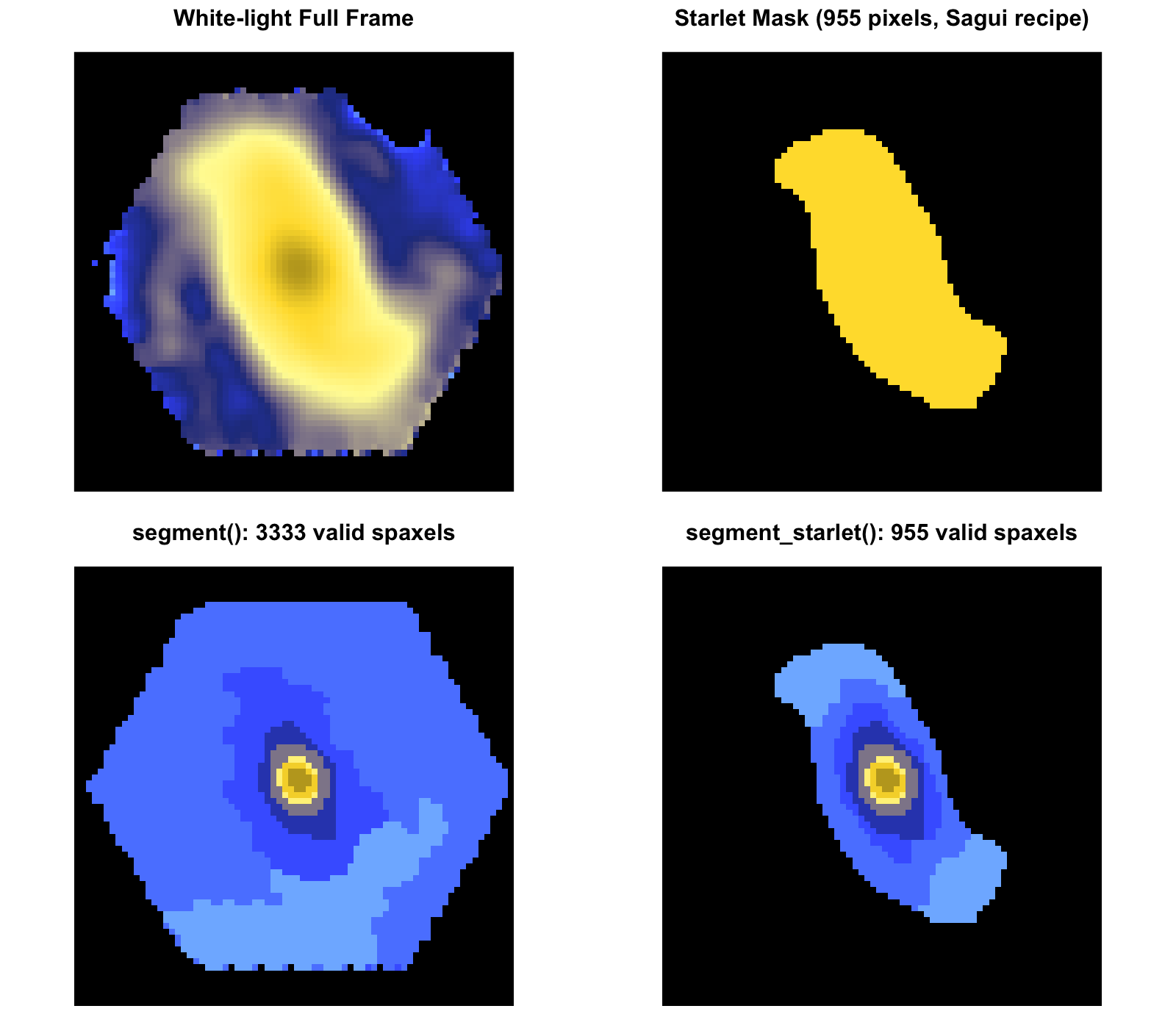
Reproducible MaNGA Examples
These panels were generated with the current public API on full MaNGA cubes, so the website examples reflect real package outputs rather than placeholder figures.
MaNGA 8135-12701
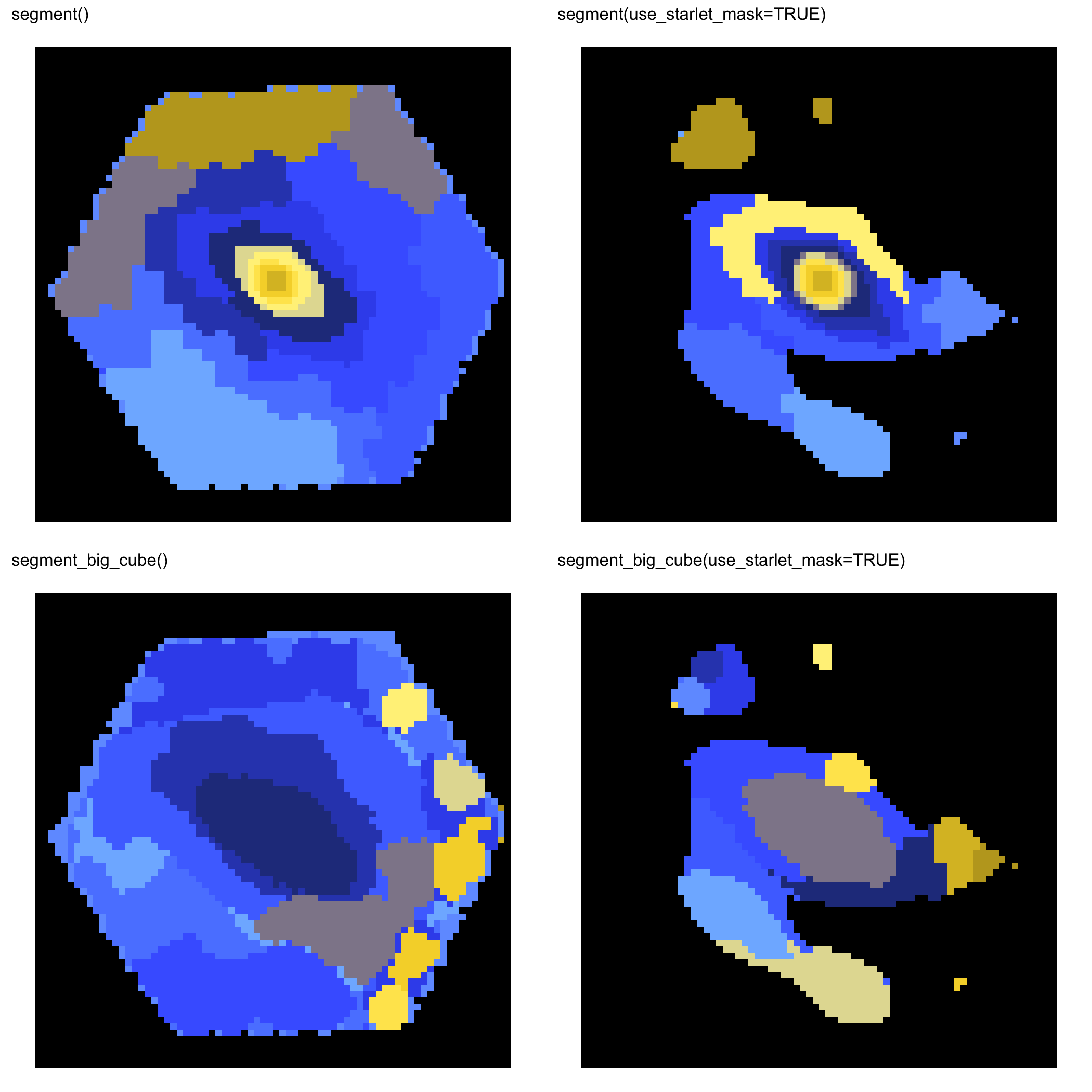
MaNGA 8443-6102
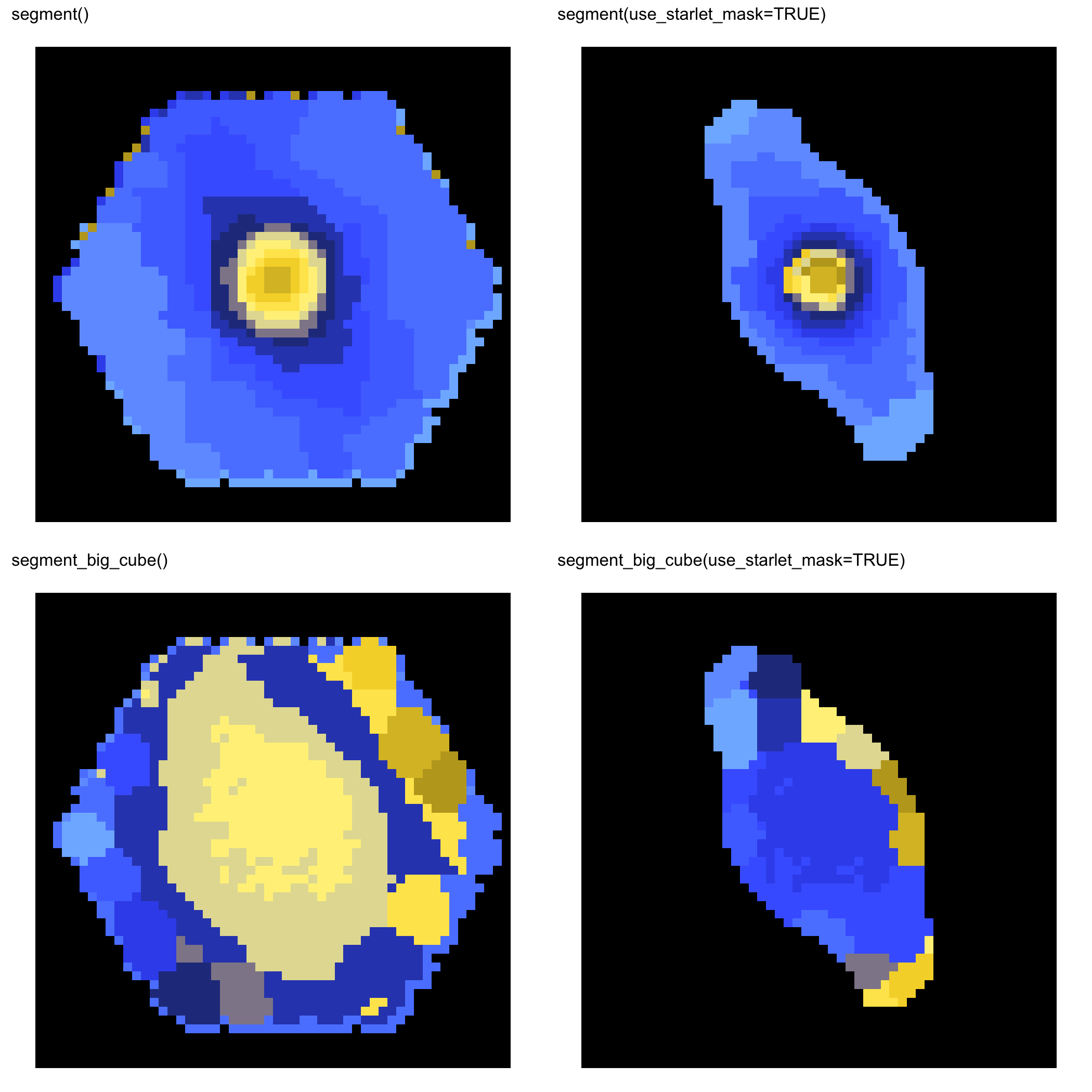
MaNGA 10224-6104
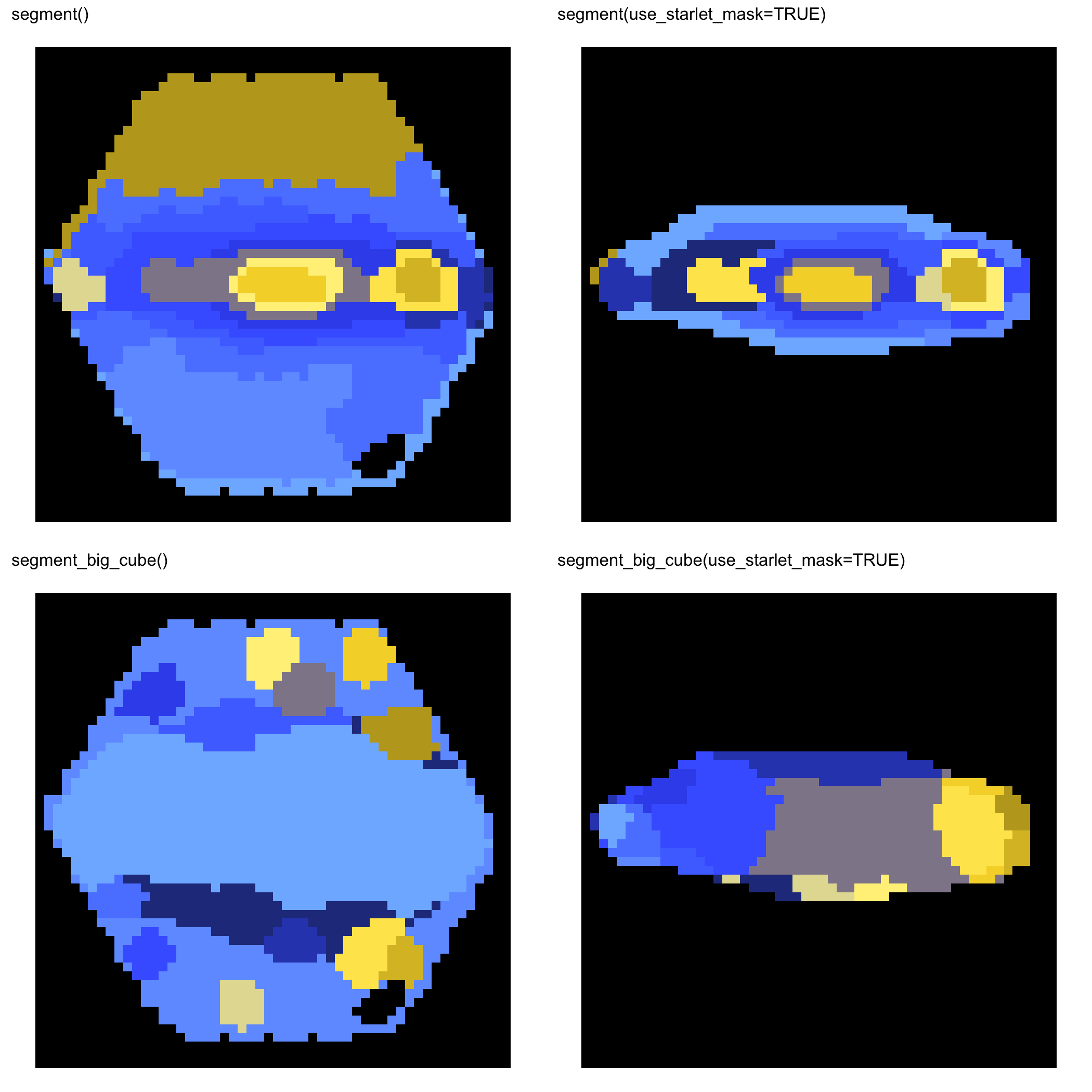

MaNGA 11749-12701
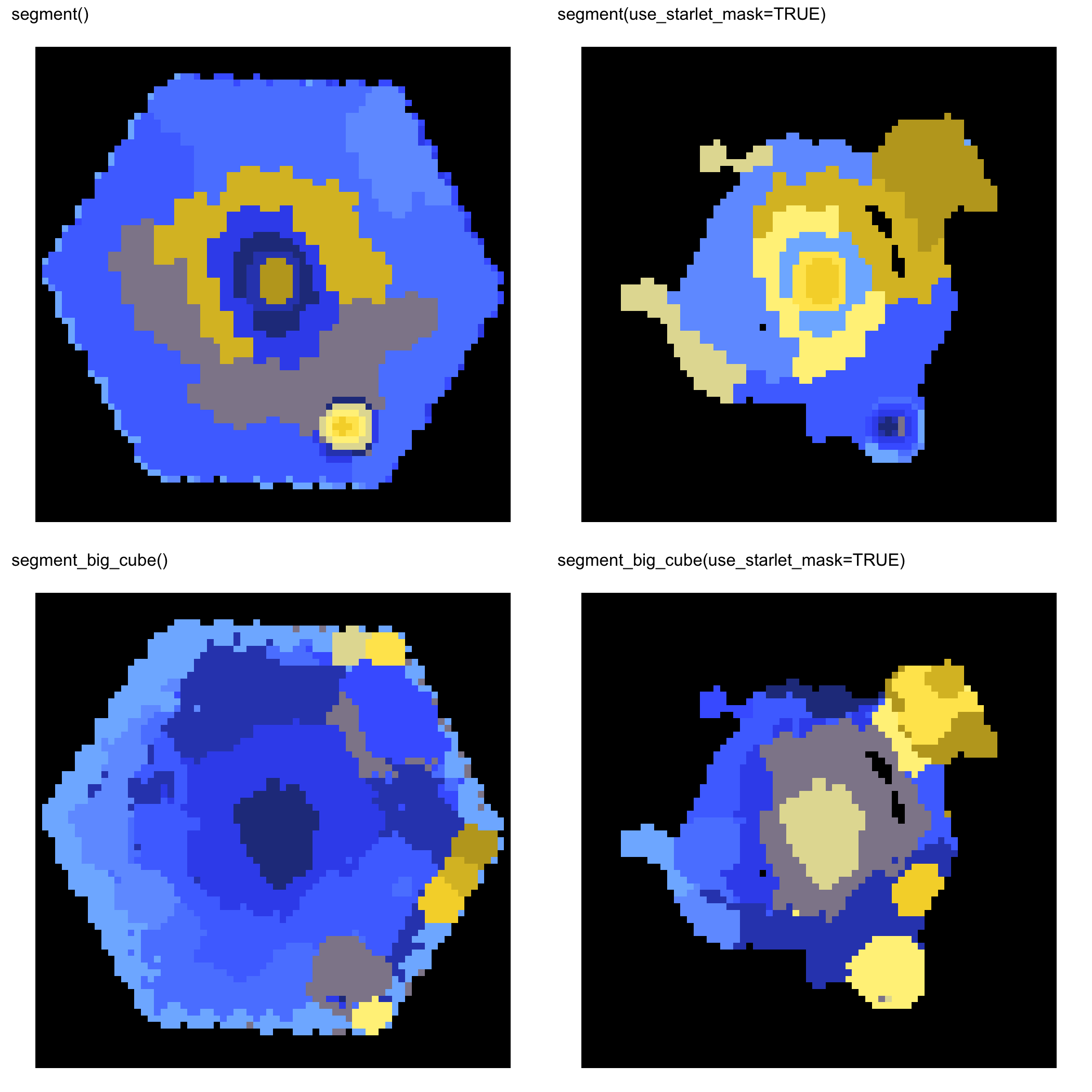

Variance-aware Summaries
Reconstructed Cubes
Attribution
If you use the Capivara code in your research, please cite the Capivara paper.
Dependencies
- torch: Optional GPU-accelerated tensor computations.
- ggplot2: Visualization.
- FITSio: Reading and handling FITS files.
- reshape2: Data manipulation.
References
- MaNGA Survey: Bundy, Kevin, et al. “Overview of the SDSS-IV MaNGA Survey: Mapping Nearby Galaxies at Apache Point Observatory.” The Astrophysical Journal 798.1 (2015): 7. DOI: 10.1088/0004-637X/798/1/7
- Capivara Code: RafaelSdeSouza/capivara
- Capivara Methodology: Souza, R. S. de, et al. (2025). CAPIVARA: A spectral-based segmentation method for IFU data cubes. Monthly Notices of the Royal Astronomical Society, 539(4), 3166–3179. https://doi.org/10.1093/mnras/staf688
- Torch in R: Paszke, Adam, et al. “PyTorch: An Imperative Style, High-Performance Deep Learning Library.” Advances in Neural Information Processing Systems 2019.



